Faculty Profile: Sebastian Behrens

Molecular Ecology for Environmental Engineering - Managing Microbial Processes That Provide Service to Society
Microorganisms are the most diverse and abundant cellular life forms on Earth. They occupy every possible metabolic niche, yet the vast majority of Earth’s bacteria cannot be analyzed using classical microbiological methods, hence they remain mysterious, sometimes referred to as the “dark matter of biology.” Microbiologists have only recently become aware of this “biological dark matter” through modern DNA sequencing surveys based on conserved marker genes (chiefly small subunit ribosomal RNA; SSU rRNA) or through random shotgun sequencing (metagenomics). Analyzing the genetic makeup of “dark matter” microorganisms could unlock a vast repertoire of new and useful metabolic functions and chemical compounds.
Microbial Ecology
Microbial ecology is the scientific discipline that aims to understand microorganisms, the microbial communities in which they live (including Bacteria, Archaea, and unicellular Eukaryotes) and how the communities interact with abiotic environments. Microbial ecologists strive to answer four central questions:
Who is in the community? This questions addresses the number and identity (phylogenetic classification) of the microorganisms present in a community.
What are the community members capable of? That is, what reactions and transformations can the community carry out that have an impact on the community’s environment? These metabolic capabilities are referred to as the community’s functional potential.
What is the community actually doing? The reactions and transformations that the community members are actually performing are called the community’s activity.
How do the microorganisms interact among themselves and with their environment? Interactions can be described based on spatial organization (who is close to whom) and chemical compounds (metabolites) that the microorganisms exchange. Understanding the complex interplay of microorganisms within a community and with their abiotic environment is the ultimate goal of microbial ecology.
Microbial ecologists made significant strides toward answering these questions with the advent of molecular biology tools. These modern molecular tools for determining the diversity within microbial communities are becoming increasingly accessible. Now a major challenge for microbial ecologists is linking the identities and structures of natural microbial communities with their functions.
"Microorganisms are absolutely fascinating. I am motivated every day to learn more about the hidden world of tiny cells that run our planet."
The first fundamental question of microbial ecology, “Who is out there?” has been addressed through use of selective and reliable amplification of conserved marker genes using the polymerase chain reaction (PCR) and hybridization with fluorescent labeled DNA oligonucleotide probes. These tools made it possible to directly interrogate the structural composition of entire microbial communities. The small-subunit ribosomal RNA (SSU rRNA, also known as 16S rRNA of bacteria and archaea) is the most widely used target gene for classification and phylogenetic inference in these applications.
Targeting genes that encode specific metabolic functions of microorganisms help answer questions about the functional capability of microorganisms. For example, amplifying and detecting specific metabolic genes from microbial genomes provides information on their genetic potential, addressing the second major question.
Messenger RNA (mRNA) reveals which genes are being expressed and, therefore, which functional proteins are likely to be formed. Information on the expressed phenotypic potential helps answer the third question about the community’s actual activity.
Assessing RNA targets by using microscopic visualization, such as with fluorescence in situ hybridization (FISH), adds information on spatial organization. Microbial ecologists have also begun to randomly sequence DNA from environmental microbial communities and it is now possible to reconstruct entire genomes of uncultured microbial community members. In addition, high-resolution imaging mass spectrometry (nano secondary ion mass spectrometry, nanoSIMS), coupled with FISH (i.e., FISH SIMS) and stable-isotope probing, make it possible to directly link a specific metabolic activity of a population of individual cells in a complex community with their phylogenetic identification, even if the encoding functional genes are unknown and the respective microorganisms cannot be grown in pure culture in the laboratory.
Application-oriented research for environmental management
Microbial Ecology builds the foundation of Environmental Biotechnology. Microbial Ecologists increase understanding of how microbial communities work. Environmental Biotechnologists manage microbial communities to provide services to society. Environmental Biotechnologists aim to manage or engineer system conditions that lead to robust communities that can provide services, such as contaminant clean up: removing contaminants from water, wastewater, sludge, sediment, or soil; resource recovery: capturing valuable products like energy, nutrients, metals, or water; sensing contaminants or pathogens in the environment, or perhaps, in humans; and protecting the public from exposure to dangerous pathogens. These services are essential to keep society safe, sustainable, and secure. Properly managed microbial communities can provide these services reliably, continuously, economically, and without creating other hazards.
Behrens and his research group strive to gain a fundamental understanding of the structure and functioning of microbial communities to better manage processes such as biological wastewater treatment, contaminant clean up, or resource recovery. Their research focuses on urgent environmental issues, including food, water, energy, waste management, and climate change.
They combine interest in fundamental microbial processes with application oriented research and focus on the basic principles of interaction between microbes, minerals, and organic matter (aka biogeochemical interfaces). A better understanding of biogeochemical processes at environmental interfaces is crucial for predicting and managing the toxicity, mobility, bioavailability, and biodegradation of organic and inorganic contaminants in natural and engineered ecosystems.
An important consideration has been the role of spatial and temporal scale. Microorganisms are generally only a few micrometers in size. Even microbial aggregates are only occasionally larger than 1 mm. However, the technological processes in which microbial communities are used (for example, wastewater treatment plants) are much larger. A bench-scale bioreactor has dimensions in centimeters, and full-scale systems measure in meters or tens of meters. One of the major challenges is relating processes that happen at the micro scale to what happens at the much larger scale of ecosystems or bioreactors.
One of Behrens’ most recent interests is the use of biochar, which combines many of his fundamental research interests with large scale environmental management strategies and new sustainable solutions for various environmental challenges. Biochar is a product of thermal degradation of organic materials in the absence of air (pyrolysis). Recently, Behrens got interested in the concept of applying biochar (aka charcoal or pyrogenic carbon) to mitigate environmental issues. Soil biochar amendment can reduce leaching of nutrients from soil, reduce the bioavailability of environmental contaminants, sequester carbon, reduce greenhouse gas emissions, and enhance crop productivity. Biochar has been applied to mitigation of large scale environmental management issues such as soil improvement (increased productivity or reduced pollution), waste management, bioremediation, and climate change mitigation. More recently ‘engineered’ or designed biochars, which are chemically modified to optimize their properties for specific purposes, have been used for metal sorption and nitrate/ phosphate capture.
Effective application of biochar requires a mechanistic understanding of microscale processes involving complex microbial communities and their interactions with minerals, natural organic matter, and the char. Also required is a comprehensive understanding of the complementary and synergistic social, economic, and life-cycle aspects of various biochar systems. Any potential solution to environmental problems based on a mechanistic understanding of biochar-microbe-mineral interactions with contaminants of environmental concern requires a whole-life-cycle assessment of the biochar application systems, for example, removal of sulfate and heavy metal from mine waters or development of a nitrate/phosphate slow-release fertilizer that prevents nutrient leaching.
Although biochar is just one of many potential solutions, it has great potential for addressing future challenges. Biochar systems can address a variety of objectives, can operate on different scales, and can be very different from each other. So adoption of biochar-based strategies could occur in multiple sectors, for example, agriculture, wastewater treatment, energy production, or climate change mitigation.
Biochar research and sustainable solutions will require interdisciplinary research teams and cross-sector collaborations with partners from industry, investors, policy makers, and practitioners. Working as a biologist in an engineering department provides Behrens with the interdisciplinary environment and the complementary expertise to take on the challenges associated with process-oriented management of microbial communities that help to clean toxic pollutants from our precious water resources and natural ecosystems.
Method Development: Exploring Diversity with Single-cell Genomics
METAGENOMICS (the study of genetic material found in natural environmental samples) has enhanced our understanding of the great diversity of microorganisms. However, when looking at a complex microbial community, the genomes of individual species often cannot be assembled, and therefore, the direct link between a cell’s identification (phylogenetic classification) and its metabolic capabilities is often lost making it difficult to understand who is doing what, where and when.
For this reason, many complete genome sequences have been obtained mostly from cultured microorganisms. Single-cell genomics conserves the genetic information that links a cell’s identification with its metabolic capabilities. Making this linked information accessible through large-scale, high-throughput studies does allow researchers to classify microbial genomes and help to identify how their genetic potential relates to observed processes in environmental samples.
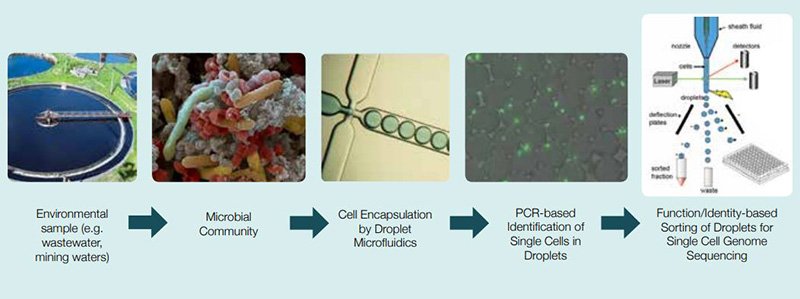
In Behrens’ laboratory, they develop new techniques to isolate and characterize single cells from environmental samples, including groundwater water, fresh water, wastewater and sediments. A new technique is based on droplet microfluidics and fluorescence activated cell sorting (FACS) using multiple displacement amplification to obtain single amplified genomes (SAGs).
Further development of single-cell technologies and integration with functional cell screening assay will make this technique a robust tool for assessing the metabolic diversity of microbial communities in natural and engineered (eco)systems.
"We live in a microbial world."

“During my graduate studies, I discovered that microbes are everywhere around us, in and on our bodies, responsible for our health, the very engines that facilitate life on Earth. It might surprise you to learn that the number of bacterial cells living in and on our bodies exceeds the number of human cells by up to an order of magnitude. Yet these human-associated bacteria, as well as most bacteria in the environment are, to a large extent, unknown and uncharted.
Exploring the tremendous genetic and functional diversity of microorganisms is like walking into an unknown forest. I discover new forms of life, observe their behaviors and life styles, and discover how the organisms interact with each other and with other living things, and I see how they affect their ecosystems.
One big challenge is that microbial “forests” cannot be seen by the naked eye: bacteria measure just a thousandth of a millimeter. So my explorations require a lot of creativity. I have to think of ways to identify and observe behaviors of bacteria in their natural environments. Modern molecular biological tools and imaging techniques help make this exploration possible.
It is regrettable that most people only recognize the negative side of bacteria, see them only as infectious agents of transmittable diseases. The majority of bacteria are good. They help in agriculture and food processing, and they play important roles for wastewater treatment and for remediation of contaminated soils and waters. They also fulfill a multitude of important functions in the human body.
Microorganisms are absolutely fascinating. I am motivated every day to learn more about the hidden world of tiny cells that run our planet.”